Published on: January 13, 2023
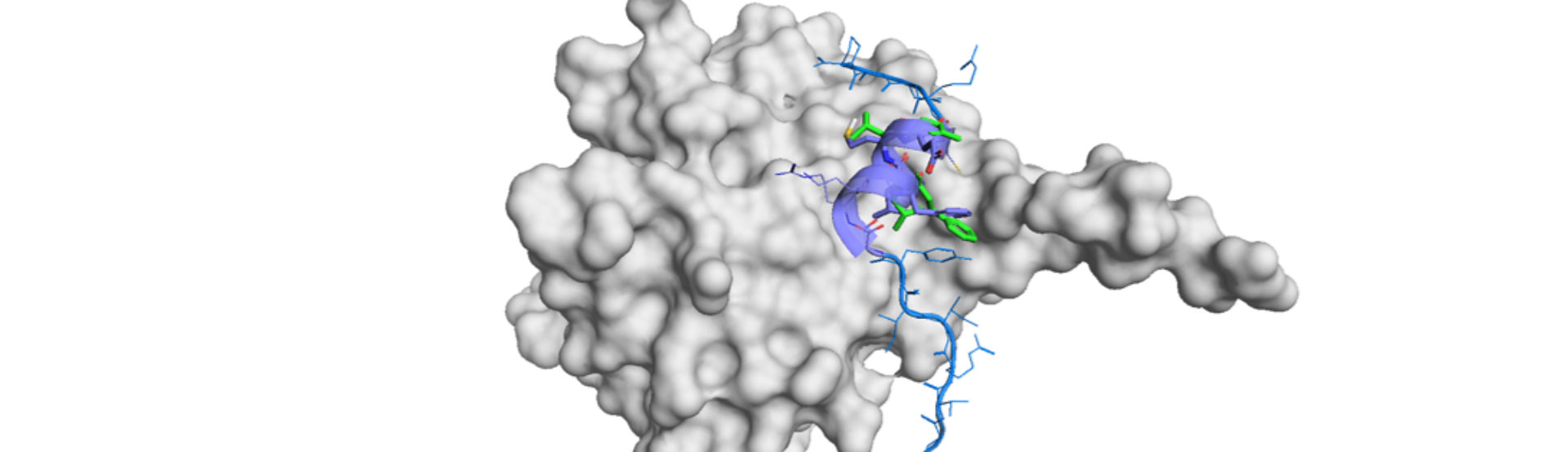
The paper published in the special issue of Pharmaceuticals journal dedicated to “Small Molecules Targeting Protein-Protein Interactions (PPIs): Current Strategies for the Development of New Drugs” presents a new method to generate small-molecules PPI inhibitors using only a sequence of a binding peptide. Potent and active PPI inhibitor peptides can be readily generated by various in vitro methods, but they are difficult to develop as drugs due to poor pharmacokinetic properties in vivo. On the other hand, high throughput screening of small molecule libraries has not been a successful approach for the discovery of PPI inhibitors. Thus, technologies that could utilize binding peptides as a starting point to design small molecules would be especially useful for the discovery and development of drugs against traditionally “undruggable” targets such as protein-protein interaction inhibitors. In this study, scientist from PRISM Biolab have utilized PepMetics technology to generate small molecules that mimic a cyclic peptide inhibitor of CTLA-4 and B7-1 PPI. By only using a sequence the cyclic peptide, they have designed a series of PepMetic molecules that mimic sequential 3-5 amino acid stretches of the peptide and have identified a molecule which upon limited optimization yielded a low micromolar PPI inhibitor. This PepMetics technology-based strategy is quick, does not require X-ray structures of a peptide bound to the target protein, and can be used to derive small molecule inhibitors from any type of a peptide. The study was done in collaboration with Gene Frontier Corporation1 and Institute for Theoretical Medicine, Inc.2
Pharmaceuticals 2022, 15(12), 1506